A newly came upon molecule disrupts most cancers cells’ talent to fix DNA via triggering the breakdown of key proteins.
Cancer cells can live on via repairing harm to their DNA—even harm that will usually be deadly. A key protection mechanism is homologous recombination, a extremely correct restore procedure that fixes damaged DNA the usage of proteins reminiscent of RAD51 and CHK1. Remedies like PARP inhibitors goal this weak spot, however many tumors in the end get better their restore talent and transform resistant.
A analysis crew led via Kyungjae Myung on the Heart for Genomic Integrity throughout the Institute for Fundamental Science (IBS), operating with Joo-Yong Lee (Chungnam College), has known a brand new manner to triumph over this resistance. Their findings display that most cancers cells may also be made inclined once more, now not via converting genetic mutations, however via disrupting the stableness of the DNA restore equipment itself.
DNA restore proteins are repeatedly regulated to care for a stability between solving harm and retaining genome steadiness. The researchers discovered that this stability may also be deliberately disturbed.
Discovery of a Key Molecule
The usage of a cell-based screening strategy to find out about replication rigidity responses, the crew known a small molecule known as UNI418. When implemented to most cancers cells, UNI418 sharply lowered ranges of key restore proteins, together with RAD51 and CHK1. As those proteins declined, the cells misplaced their talent to fix DNA successfully.
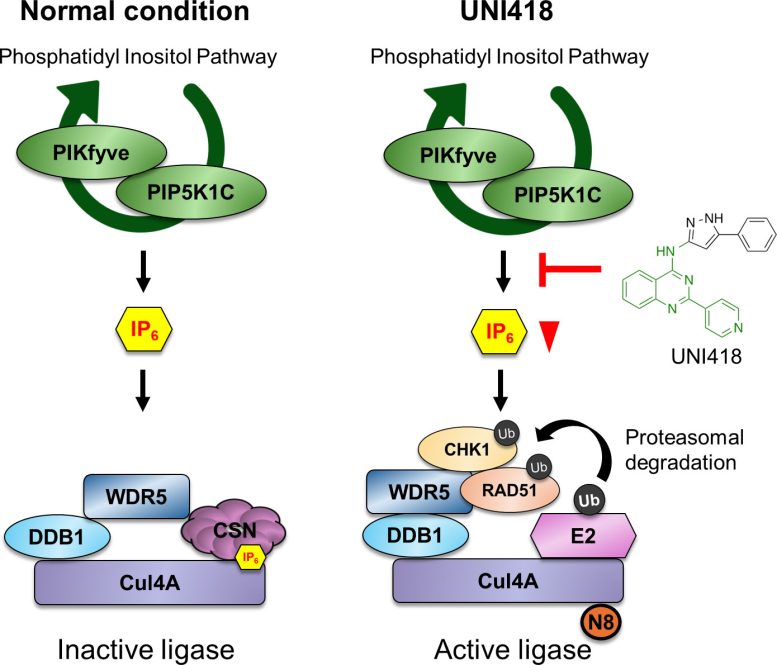
Additional investigation confirmed that UNI418 turns on a protein degradation device referred to as the Cul4A ubiquitin ligase complicated, which marks particular proteins for destruction. This procedure successfully breaks down the DNA restore device from inside.
Co-corresponding writer Professor Joo-Yong Lee mentioned, “We known a mechanism wherein key DNA restore proteins are actively degraded within the mobile. This gives a brand new technique to keep an eye on homologous recombination past genetic mutations.”
The Position of Mobile Metabolism
The researchers then explored how this degradation procedure is precipitated. UNI418 interferes with a signaling pathway curious about inositol phosphate metabolism, decreasing ranges of a molecule known as IP6. Underneath standard prerequisites, IP6 suppresses Cul4A job. When IP6 ranges drop, this suppression is got rid of, permitting the degradation device to turn on.
Cul4A, together with its adaptor protein WDR5, then objectives DNA restore proteins reminiscent of RAD51 for destruction, shutting down homologous recombination.
This creates a state very similar to DNA restore deficiency, even in most cancers cells that had up to now regained restore serve as. This impact is particularly related for overcoming resistance to PARP inhibitors, a significant problem in most cancers medication.
The crew examined whether or not this technique may beef up remedy results. In numerous cell-based experiments, UNI418 made most cancers cells extra delicate to PARP inhibitors. It additionally labored in cells that had already evolved resistance, restoring their reaction to medication.
Co-corresponding writer Kyungjae Myung added, “By way of weakening the DNA restore device, we will be able to re-sensitize tumors that experience transform immune to current remedies. This means a brand new technique for increasing the effectiveness of PARP inhibitors.”
The findings had been additionally showed in animal fashions. In tumor xenograft experiments, UNI418 lowered tumor expansion, particularly when blended with the PARP inhibitor Olaparib. This impact used to be seen even in fashions designed to mirror treatment-resistant cancers.
Broader Implications for Cancer Remedy
The findings had been additionally showed in animal fashions. In tumor xenograft experiments, UNI418 lowered tumor expansion, particularly when blended with the PARP inhibitor Olaparib. This impact used to be seen even in fashions designed to mirror treatment-resistant cancers.
Those effects display that most cancers cells nonetheless rely on DNA restore programs even after changing into resistant, and that disrupting protein steadiness can disclose this dependence.
The find out about additionally highlights a up to now unknown hyperlink between cell metabolism and DNA restore. By way of connecting IP6 signaling with the Cul4A-driven degradation pathway, the researchers known a brand new mechanism that is helping keep an eye on genome steadiness.
Co-corresponding writer Kyungjae Myung remarked, “This find out about demonstrates that controlling the stableness of DNA restore proteins can without delay have an effect on most cancers mobile survival. It additionally highlights a brand new healing route for overcoming drug resistance.”
Total, the findings recommend that drug-resistant cancers may also be made inclined once more via breaking down the programs they use to fix DNA slightly than changing their genes. Even if UNI418 would require additional building, the mechanism described on this find out about provides a robust basis for long term mixture remedies.
Reference: “Concentrated on IP6 signaling to destabilize homologous recombination proteins to triumph over PARP inhibitor resistance” via Seon-gyeong Lee, Yuri Web optimization, Seula Jeong, Yuheon Chung, Sukyeong Kong, Minyoung Kim, Joon Ho Rhlee, Sihyeon Um, Bijoy P. Mathew, Saikat Maiti, Malleswara Rao Kuram, Mohamed Ahmed Abozeid, Areum Park, Ji-Na Yoo, Keon Woo Khim, Kyuwon Son, Enkhzul Amarsanaa, Kyunghan Kim, Sehoon Hong, Jiyeon Choi, In Bae Park, Eun A. Lee, Ji Hwan Jeon, Jun Hong Park, Joo Seok Han, Chan Younger Park, Seyun Kim, Jang Hyun Choi, Sung You Hong, Min-Duk Web optimization, Hyuk Lee, Joo-Yong Lee and Kyungjae Myung, 4 April 2026, Nature Communications.
DOI: 10.1038/s41467-026-71421-z
The find out about used to be funded via the Institute for Fundamental Science.
By no means leave out a leap forward: Join the SciTechDaily newsletter.
Apply us on Google and Google News.